
TMIC and the Metabolomics Society
Issue 40 - December 2014
CONTENTS:
Online version of this newsletter:
http://www.metabonews.ca/Dec2014/MetaboNews_Dec2014.htm
 |
| Published
in partnership between TMIC and the Metabolomics Society Issue 40 - December 2014 |
|
CONTENTS: |
|
|
 |
Metabolomics Society News |
| |
Metabolomics Spotlight
|
 |
 |
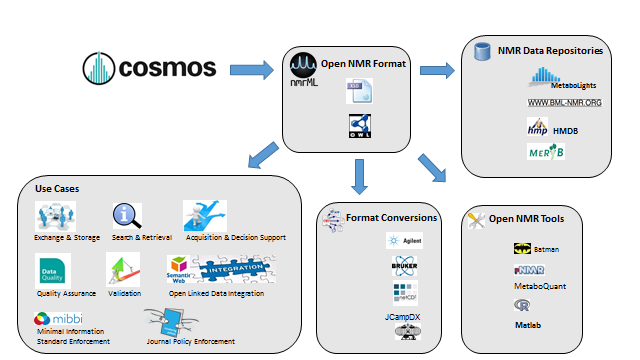
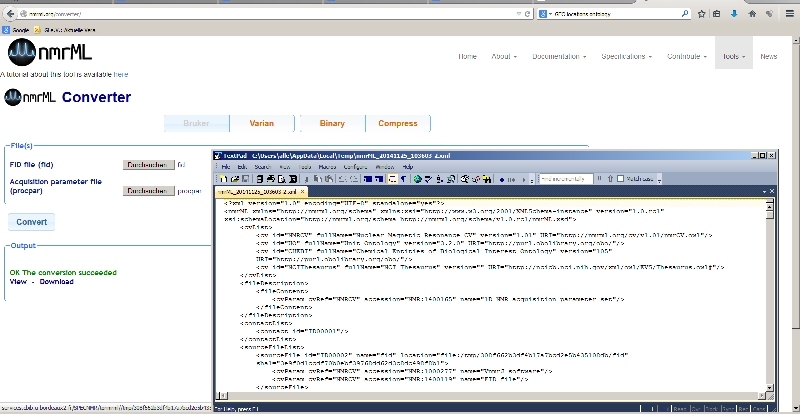
| |
MetaboInterviews
|
| Vice President, Business and Product Development, Human Metabolome Technologies, Boston, USA |
|
 |
Biography I received my Bachelor’s degree in 1975 from University of Virginia in Geochemistry and a PhD in 1980 from University of Virginia in Chemistry developing mass spectrometric sequencing of proteins. From 1980 to 1984, I completed a post-doctoral fellowship at the FDA Bureau of Biologics and Biophysics, studying sequencing of inflammatory cytokines and natriuretic factors in blood. I worked at Abbott Laboratories in Chicago from 1984 to 2002, and led a large lab analyzing small molecules, metabolites, and proteins. I worked on measuring metabolites using MALDI-TOF and experimented with capillary electrophoresis mass spectrometry (CE-MS) for proteins and small molecules, using MALDI-FTMS and LC-MS/MS. I moved to Biogen Idec from 2002 to 2012 and led a large lab analyzing small molecules, proteins, and antibodies. I built proteomic and metabolic platforms at both Abbott and Biogen Idec. Over the course of my career, I have worked on disease and drug response biomarkers in several therapeutic areas and believe in a systems biology approach using innovative data mining tools with algorithm generation. Currently I am VP of Business and Product Development at Human Metabolome Technologies. Our company has been delivering quantitative and non-quantitative CE-MS metabolic profiles for many years mainly in Japan. Metabolomics and metabolite profiling have experienced significant growth in the US and Europe over the past several years covering many areas of science. Since HMT has developed an extensive metabolite database over the past 10 years, serving many areas from agriculture to pharmaceuticals, the time has come to bring HMT metabolomics to a growing US market. |
 |
Metabolomics
Current Contents
|
 |
MetaboNews |
| 19
Nov 2014 |
Patients to benefit from new £5m
MRC Regional Phenome Centre at the
University of Birmingham, UK
The University of Birmingham, UK, has announced plans to establish a £5m MRC Regional Phenome Centre (RPC), which will help to improve the health of patients through novel research and technology. The centre will complement and work closely with the MRC-NIHR National Phenome Centre in London. It will help scientists to better detect the onset of several diseases and to develop more effective treatments, focusing primarily on patients with immune-mediated inflammatory diseases and blood cancers. Significant improvements in patient health, as well as cost savings to the UK’s National Health Service, are anticipated through early detection and tailoring of treatment. The centre will be led by Professor Mark Viant, Professor Ulrich Guenther and Dr Warwick Dunn. Building on existing metabolomics technologies in the Schools of Biosciences and Cancer Sciences at the University of Birmingham, the MRC award will fund ten additional mass spectrometers and NMR spectrometers to greatly enhance the number of samples that can be investigated. The centre will soon be recruiting six full-time metabolomics scientists, applying LC-MS, NMR spectroscopy and computational biology. The first of these six posts, the Operational Head of Mass Spectrometry and Bioinformatics for the MRC RPC, is currently being advertised (see the Metabolomics Jobs section below). The metabolomics research at Birmingham and the new centre benefit from an existing Technology Alliance Partnership between the University of Birmingham and Thermo Fisher Scientific for advancing life sciences mass spectrometry, and a technology collaboration between the University and Beckman Coulter for advancing automation solutions in the omics sciences. Collaborations with Bruker and Waters will further enhance the capacity and capabilities of the centre. Professor Viant said, “We are tremendously excited about building a metabolomics facility of the scale needed to significantly improve our ability to understand mechanisms of human disease and improve treatments. The centre will provide the UK with a significant advantage that we will exploit to generate new diagnostic and prognostic biomarkers of both common and rare diseases.” National Phenome Centre Director, Professor Jeremy Nicholson of Imperial College London, added, “We are pleased to be associated with the new MRC Regional Phenome Centre at Birmingham and this will form part of a wider network of national and regional centres that will harmonise with the NPC and the linked Imperial BRC Clinical Phenome Centre. The multiple benefits of network driven commonalities of approach, methodology, data mining and database generation are easy for all to see." The MRC RPC will form a cornerstone of the new £25m Institute for Translational Medicine, which will open in June 2015 and is governed through the Birmingham Health Partners, a strategic alliance between the University of Birmingham, University Hospitals Birmingham NHS Foundation Trust and Birmingham Children’s Hospital. Professor David Adams, Dean of Medicine at the University of Birmingham and Director of Birmingham Health Partners commented, “These are exciting new technologies that we anticipate with bring significant patient benefits, both locally and nationally.” About the National Phenome Centre The MRC-NIHR National Phenome Center is a partnership between Imperial College London and Kings College London with Waters Corporation and Bruker Biospin. Opened in July 2013 under the Directorship of Professor Jeremy Nicholson of Imperial College (http://www3.imperial.ac.uk/newsandeventspggrp/imperialcollege/newssummary/news_4-6-2013-12-3-42), the NPC provides the world's first national capability and capacity for large scale metabolic phenotyping for molecular epidemiology biomarker studies and stratified medicine programs using a multiple high field NMR and mass spectrometer cluster. For further information visit http://www1.imperial.ac.uk/phenomecentre/ |
| 5
Nov 2014 |
Steno opens new metabolomics
laboratory
Steno Diabetes Center has today officially opened its new state-of-the-art metabolomics laboratory. The goal of the new technology is bringing Steno at the forefront of personalized diabetes treatment. "The overall goal is to bring metabolomics technology into the clinic," says Matej Orešič, principal investigator of Systems Medicine at the Steno Diabetes Center. "The new technology will enable us to predict the development of the disease, which can guide health professionals in relation to establishing personalized prevention and treatment of diabetes." Metabolomics is a field of research that examines metabolites in fluids, tissues and cells. Metabolites such as glucose and cholesterol, have been extensively studied in diabetes research, while the current metabolomics technology makes it possible to examine thousands of metabolites simultaneously. Until now, metabolomics technology primarily been used in research projects, but has now become so developed that it can be used in clinical practice. "Based on a metabolomics analysis, we will hopefully be able to provide a profile of the patient and predict the risk of diabetes and diabetes complications, and the patient's response to a specific therapy. The challenge is to translate the detailed information from more than 1000 metabolites into useful information for clinical practice. It requires much training and is an interesting and relevant area to develop, "says Tuulia Hyötyläinen, head of the new metabolomics laboratory. Steno laboratory contains several instruments that provide detailed analyzes of polar metabolites and lipids, as well as quantitative analyzes of selected groups of metabolites. The new equipment is already in use in several research projects are underway in Systems Medicine research group has recently been established at Steno. The plan for metabolomics laboratory is to start with research projects and so involve patients from Steno's clinic in 3-4 months when all clinical validations are in place. "I want to congratulate Matej Orešič and Tuulia Hyötyläinen to have established their laboratory so soon after their employment at Steno," says Managing. Director John Nolan. "The new metabolomics and lipidomics laboratory is a significant step forward for the Steno Diabetes Center and will strengthen our capacity for translational research in the early development of diabetes and prevention. We look forward to establishing new collaborations with colleagues in Denmark and internationally. " Steno Diabetes Center is one of the first diabetes clinics in the world where one implements metabolomics in clinical practice. Many partners from the Danish diabetes research community attended the opening, which took place at the Steno Diabetes Center's birthday, established November 5, 1932. Source: Press Release translated into English (Google Translate) Original Press Release (in Danish) |
Metabolomics Events
|
| 3-5 Dec 2014 |
2nd
Australian Lipid Meeting While the first meeting focused on lipidomics we have expanded the scope for the second meeting to cover all aspects of lipid research. Planned topics include:
Registration
|
| 11 Dec 2014 |
London Biological Mass
Spectrometry Discussion Group The meeting will be held at Cancer
Research UK, 44 Lincoln's Inn Fields,
London, WC2A 3LY. The nearest
tubes are Holborn or Chancery Lane. To see the agenda for this event, click here. |
| 11-13 Dec 2014 |
2nd ICAN (Institute of
Cardiometabolism and Nutrition) Conference
Series 2014 Further Information: http://ican-series.com/ For further details, visit the conference
website. |
| 9-12 Feb 2015 |
3rd Annual Practical
Applications of NMR in Industry Conference
(PANIC) The PANIC meeting is intended to address topics that occur daily in industrial, government, and academic research laboratories whose primary task entails the application of NMR to a diverse set of analytical problems. Topics include quantitation, molecular structure characterization, trace component and mixture analysis, and product support for a variety of materials that include small molecules, polymers, heterogeneous mixtures, natural products, biosimilars, polysaccharides, and proteins. The conference provides in-depth discussions of the “nuts and bolts” of basic NMR experiments that accent the underlying best-practices developed to address these everyday problems. It also explores the regulatory aspects of the applications of these experiments. Greater insights will be provided into the less frequently explored techniques used in NMR such as quantitation, chemometrics, automation, relaxometry, and at/on line instrumentation. Attendees will have the opportunity to learn about and discuss applications of NMR to polymers, petroleum, food, agriculture, and nutritional supplements that are rarely discussed at other NMR meetings. Ample opportunity will be given to Network with people who are experts in inventing NMR experiments to solve real world problems in a timely and efficient manner that is demanded by working in the venue of product delivery. For more details, visit http://www.panicnmr.com. |
| 16-20 Feb 2015 |
EMBO Practical Course on
Metabolomics Bioinformatics for Life
Scientists Organizers
Participation: Open application
with selection |
| 28 Mar to 1 Apr 2015 |
Plant Metabolism Session
at the ASBMB Annual Meeting |
| 29 Jun to 2 Jul 2015 |
Metabolomics 2015: 11th
Annual Conference of the International
Metabolomics Society You are invited to join us for Metabolomics
2015, the official annual meeting
of the Metabolomics Society. For further details, visit http://metabolomics2015.org. |
Metabolomics Jobs |
This is a resource for
advertising positions in metabolomics. If you have a
job you would like posted in this newsletter, please
email Ian Forsythe (metabolomics.innovation@gmail.com).
Job postings will be carried for a maximum of
4 issues (8 weeks) unless the position is filled prior
to that date.
Jobs
Offered
| Job Title | Employer | Location | Posted | Closes | Source |
|---|---|---|---|---|---|
| Research
Fellow (Parasitology/Metabolomics) |
Monash
University |
Parkville, Victoria, Australia | 27-Nov-2014 |
18-Dec-2014 |
SEEK |
| Postdoctoral
Researcher in Metabolomics of Cancer |
Karolinska
Institutet |
Stockholm, Sweden | 27-Nov-2014 |
31-Jan-2015 |
Karolinska
Institutet |
| Operational Head of Mass Spectrometry and Bioinformatics at the MRC Regional Phenome Centre | University of Birmingham | Birmingham, UK | 19-Nov-2014 |
9-Dec-2014 |
University
of Birmingham |
| Masters
Student - The Science of Adductomics:
Application to Gestational Diabetes |
Liggins
Institute, University of Auckland |
Auckland, New Zealand | 14-Oct-2014 |
14-Dec-2014 |
Metabolomics
Society Jobs |
| PhD
Studentship |
International
Pregnancy Research Alliance (IPRA) |
Chongqing, China | 27-Aug-2014 |
31-Dec-2014 |
Metabolomics Society Jobs |
| Mass Spectrometry Technician | International Pregnancy Research Alliance (IPRA) | Chongqing, China | 27-Aug-2014 | 31-Dec-2014 | Metabolomics Society Jobs |
Equipment
|
Ian
J. Forsythe, M.Sc.
MetaboNewsEditor Department of
Computing Science
University of Alberta 221 Athabasca Hall Edmonton, AB, T6G 2E8, Canada Email: metabolomics.innovation@gmail.com Website: http://www.metabonews.ca LinkedIn: http://ca.linkedin.com/in/iforsythe Twitter: http://twitter.com/MetaboNews Google+: https://plus.google.com/118323357793551595134 Facebook: http://www.facebook.com/metabonews |
This newsletter
is published in partnership between The
Metabolomics Innovation Centre (TMIC, http://www.metabolomicscentre.ca/)
and the Metabolomics Society (http://www.metabolomicssociety.org)
for the benefit of the worldwide
metabolomics community.
 A single source destination for fee-for-service metabolic profiling including comprehensive metabolite identification, quantification, and analysis 
|