Theory of Chemometrics
Feature article contributed by Jeroen Jansen, Lionel Blanchet,
and Lutgarde Buydens,
Laboratory of Analytical Chemistry, Radboud University Nijmegen,
Netherlands
The Laboratory of Analytical Chemistry (
LAC) at
the Radboud University Nijmegen, headed by Prof. Lutgarde Buydens,
is one of the oldest chemometrics groups in the world.
Chemometrics is the research field interested in the analysis of
complex chemical data. Omics data are therefore logically of
particular chemometric interest and a key element in our research
portfolio. Conversely, scientific and technological developments
in metabolomics require ever-more advanced statistical methods
that answer the very complex biological research questions. Such
synergy between both fields may induce a next generation of
thought-provoking challenges that stretch current methods to the
limits and thereby spark the development of new approaches.
The field of chemometrics until recently relied heavily on two
methods: Principal Component Analysis to describe the chemistry of
complex analytical measurements, and Partial Least Squares
Regression to predict specific properties from these measurements.
Any literature search for 'analysis of metabolomics data' shows
how the love for these methods flourishes on. However, LAC aims to
ascend on these specific methods, to reach a strong
theoretical—currently lacking—basis for chemometrics. For this we
started a public-private partnership, through the Netherlands 'Top
Institute for COmprehensive Analytical Science and Technology' (
TI-COAST). In this project,
called 'Analysis of Large data sets By Enhanced Robust Techniques'
(ALBERT), several Dutch (bio)chemical companies collaborate with
chemometricians and statisticians from the Universities of
Nijmegen and Groningen to create a
pipeline. In this line
the
information quality within any metabolomics (or other
chemical) dataset can be quantitatively evaluated, based on solid
theoretical grounds (
Figure
1). This then forms the basis for the
pre-processing
to remove identified artefacts and thereby highlight the
biologically relevant data. This can then be
analysed by
PCA or PLS, but potentially by more advanced methods that are
tailor-made to answer the biological questions at hand. Also, here
the theoretical basis is maintained, because each step in the
pipeline is supported by two essential, but often overlooked
aspects:
validation and
visualisation. This
pipeline will assure peer metabolomics researchers they can both
trust and understand results from the high-quality chemometric
models at the basis of a joint publication.
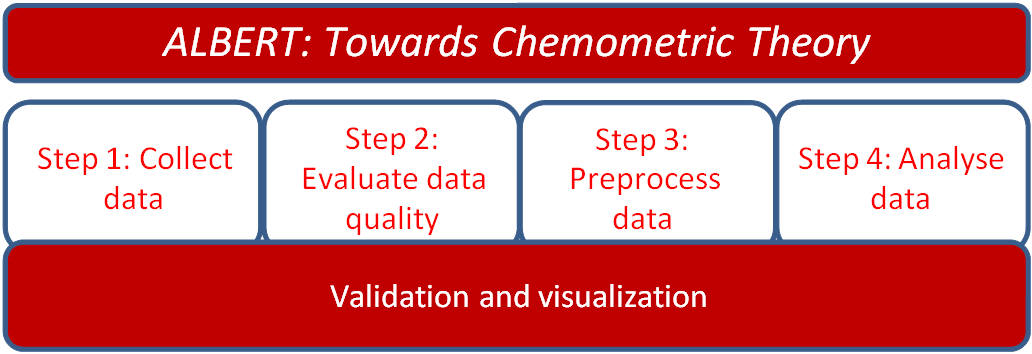 Figure 1.
Figure 1. Schematic depiction of the steps in the ALBERT
pipeline.
Our laboratory has applied this pipeline and their interconnection
in several studies. For example, we Intuitively expect that the
concentration of a compound within a particular metabolic pathway
changes proportionally to other metabolites in the same pathway.
However, in reality, these concentration changes may follow much
more complex behaviour: metabolic systems—biological problems in
general—are inherently non-linear. The chemometrician's best
friends (PCA and PLS) are unable to detect these relations in such
cases and more advanced methods are required.
Kernel
transformations are particularly promising for this, for two
reasons. Kernels can linearize almost any type of relation, which
are then open for exploration by PCA or PLS. Although kernel-based
methods are generally recognized as powerful, their use was
severely hampered by the impossibility of visualizing the
biological information (metabolites) that underlie the model,
e.g., in a 'biplot'. Our group recently solved this problem, by
implementing 'pseudo-samples' [
1]
that reconstruct the role of each metabolite in the model, even
for nonlinear biological differences. This aspect can even be
further extended to the linear and non-linear complementary
information that GC-MS and NMR generate for complex diseases like
Multiple Sclerosis (
Figure
2) [
2].
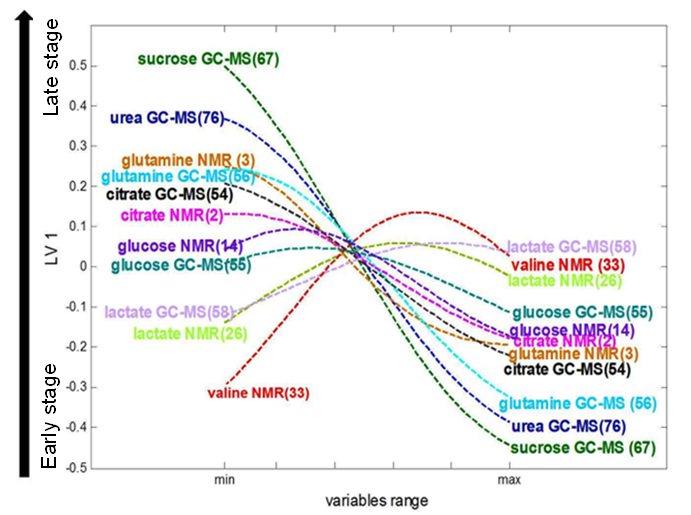 Figure 2.
Figure 2. Loading plot of pseudo-samples trajectories for
selected NMR and GC-MS variables, adapted from reference [
2].
Another
forte of chemometrics that is very useful to
metabolomics, is the strong
analytical basis that
underlies it. This is not only reflected in analytical chemistry
but also in the thought processes that lie beneath. Chemometrics
therefore forms an abstract plane, on which concepts from remote
scientific fields like psychology and marketing may
cross-fertilize with those in systems biology. A prominent field
in these social science disciplines is the study of '
individual
differences' that can be very useful to create an
'individualized metabolomic fingerprint'. In collaboration with
the
Biosystems Data Analysis
group at the University of Amsterdam, we have modeled the
ecological response of a group of plants to herbivory,
specifically the heterogeneity in this response between otherwise
completely comparable plants and the emergence of
sub-groups—chemotypes—in this response. For this we adapted the
Simultaneous Component Analysis with Individual Differences
constraints (SCA-IND) method [
3],
originally developed for assessing differences between sensory
panel members for product tests. This method provided a view on
this heterogeneity that was both ecologically and biochemically
understandable (
Figure
3). We believe this method can be applied broadly in
metabolomic studies. Specifically its application in Personalized
Health and Medicine, which focuses on the heterogeneity among
patients, seems extremely promising. This is another very clear
example of how chemometrics may facilitate cross-fertilization
between research fields, even within biology.
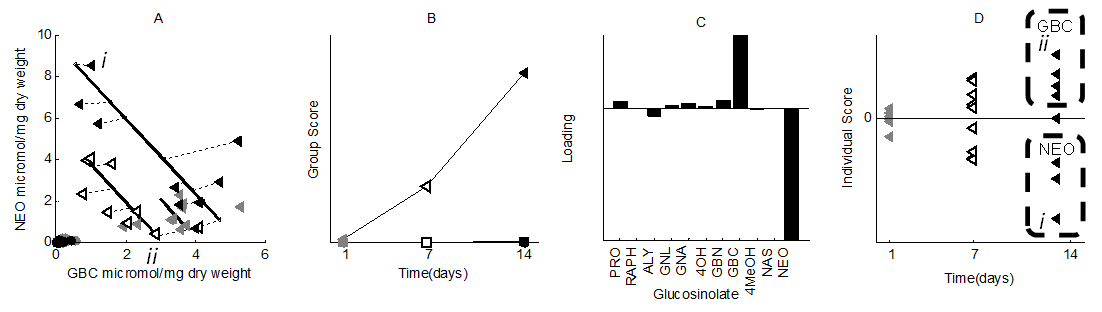 Figure 3.
Figure 3. SCA-IND model on the ecological plant response.
A. Raw data (circles: control plants, triangles: attacked plants;
grey: 1 day after attack, white: 7 days after, black: 14 days
after. A negative relation between NEO and GBC clearly emerges
during the experiment i. plant with high level of
Neoglucobrassicin (NEO), and ii. High-Glucobrassicin Plant); B.
This emergence becomes clear in the 'Group-level' scores of
attacked plants; C. Loadings show this heterogeneity mainly
associates with Glucobrassicin and Neoglucobrassicin; D. The
Individual-level scores of the model show distinct Glucobrassicin
and Neoglucobrassicin-responding plant groups, 14 days after
attack—the emergence of 'Response Chemotypes'; plants i. and ii.
from panel A are indicated.
References
- Postma, G.J., P.W.T. Krooshof, and L.M.C. Buydens, Opening
the kernel of kernel partial least squares and support vector
machines. Analytica Chimica Acta, 2011. 705(1–2): p.
123-134. [PMID:
21962355]
- Smolinska, A., et al., Interpretation and
Visualization of Non-Linear Data Fusion in Kernel Space: Study
on Metabolomic Characterization of Progression of Multiple
Sclerosis. PLoS ONE, 2012. 7(6): p. e38163. [PMID:
22715376]
- Jansen, J., et al., Individual differences in
metabolomics: individualised responses and between-metabolite
relationships. Metabolomics, 2012. 8(0): p. 94-104. [PMID:
22593723]
Please
note: If you know of any
metabolomics research programs, software, databases,
statistical methods, meetings, workshops, or training
sessions that we should feature in future issues of this
newsletter, please email Ian Forsythe at metabolomics.innovation@gmail.com.