
TMIC and the Metabolomics Society
Issue 65 - January 2017
CONTENTS:
Online version of this newsletter:
http://www.metabonews.ca/Jan2017/MetaboNews_Jan2017.htm

 |
| Published
in partnership between TMIC and the Metabolomics Society Issue 65 - January 2017 |
|
CONTENTS: |
|
|
 |
| TMIC
Services |
 |
Metabolomics Society News |
Hanne Bertram (University Aarhus, Denmark) on food metabolomicsSo visit the website for the latest information about the 2017 conference, accommodation and social activities for the Brisbane 2017 meeting, and start registering—we are looking forward to welcoming you in Brisbane!
Debra Meyer (University of Johannesburg, South Africa) on medical metabolomics
Roy Goodacre (University of Manchester, UK) on advancing the field
| |
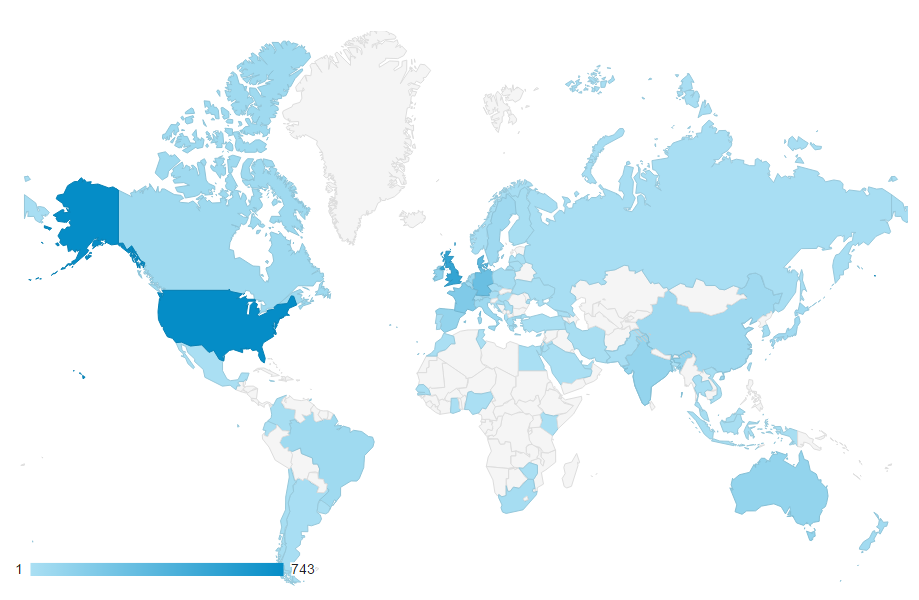
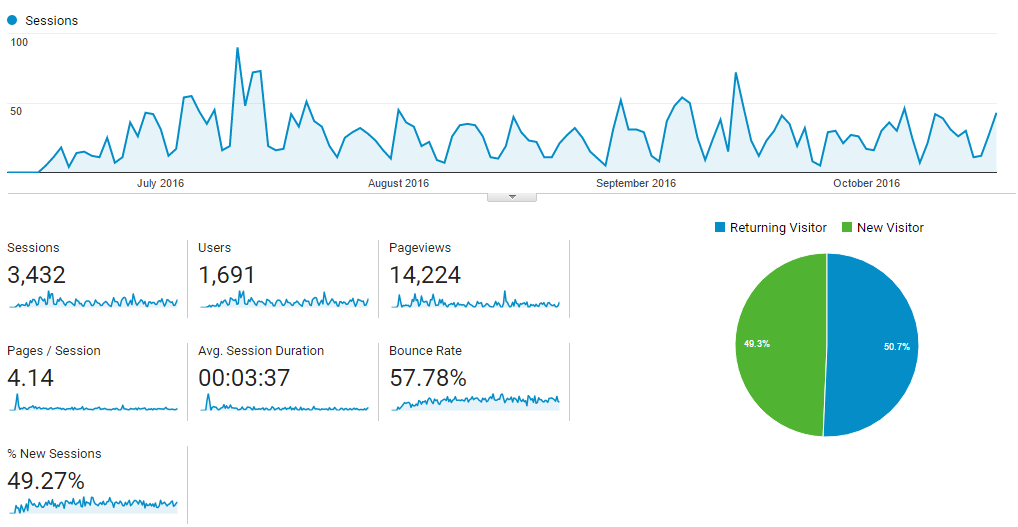
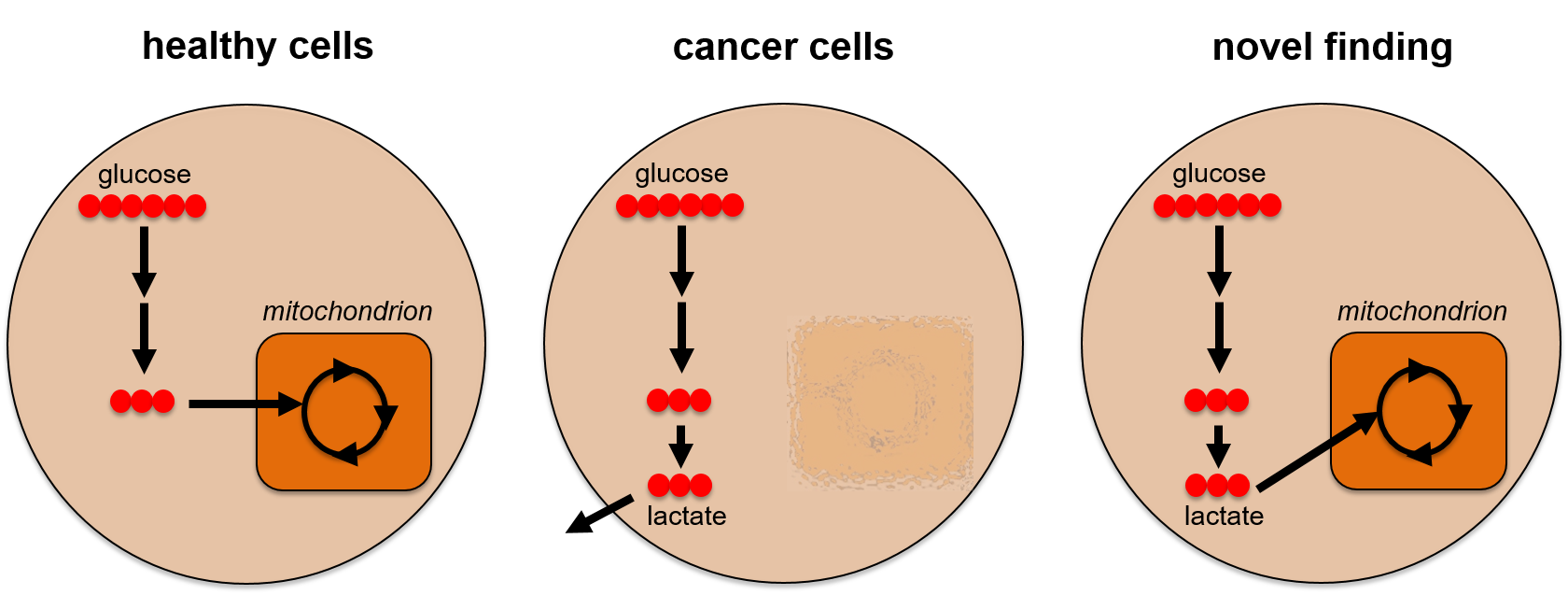
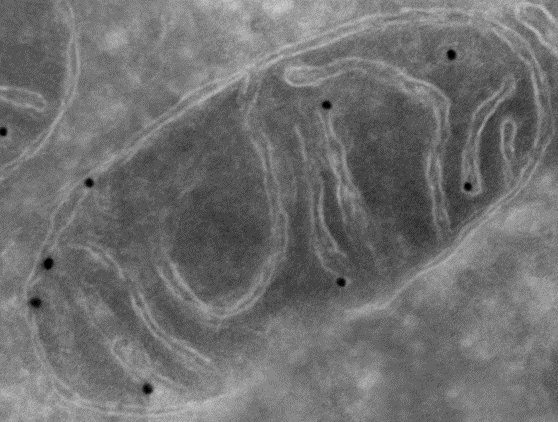
Metabolomics Spotlight
|
| |
MetaboInterview
|
| Professor in Biochemistry and
Molecular Biology at the University of Melbourne,
Australia |
|
 |
Biography Malcolm J. McConville, PhD, is Professor in Biochemistry and Molecular Biology at the University of Melbourne, Australia. He completed his PhD studies at the University of Melbourne in 1986 and subsequently held post-doctoral positions at the Walter and Eliza Hall Institute of Medical Research, Melbourne and the University of Dundee, Scotland, before returning to Melbourne in 1996. He is currently Director of the multidisciplinary Bio21 Institute of Molecular Science and Biotechnology at the University of Melbourne and head of the Metabolomics Australia node in the Institute. Malcolm's group is interested in understanding how different bacterial and protozoan pathogens survive within host cells/tissues with the view of developing more effective anti-microbial drugs. His group uses mass spectrometry-based metabolomics and stable isotope labelling approaches to dissect and model the metabolism of intracellular pathogens and host cells, and to understand the mode of action of antimicrobial drugs and pathogen resistance mechanisms. |
 |
Metabolomics Current
Contents
|
Metabolomics Events
|
| 23-27 Jan 2017 |
Hands-on NMR for Metabolic Phenotyping Venue: Imperial College London, South Kensington, London, UK This week long course aims to cover how to perform a metabolic profiling experiment, from start to finish. It will cover study design, sample preparation, NMR spectrometer set up for global profiling, 2-dimensional NMR experiments and spectral data analysis. It combines lectures and tutorial sessions to ensure a thorough understanding of the theory and practical applications. Topics covered include:
or contact Rebecca Smith (rebecca.smith2@imperial.ac.uk) for further information. |
| 8-9 Feb 2017 |
Computational Environmental Metabolomics Course Venue: Birmingham Metabolomics Training Centre, University of Birmingham, Birmingham, UK This 2-day NERC funded Advanced Training Short Course will provide a theoretical overview and hands-on training in the processing and analysis of metabolomics data. You will study specific case studies from the environmental sciences, learning the step-by-step processes that are applied in the data analysis pipeline. The course will be delivered using a combination of lectures and computer workshops, and time will be dedicated to answering questions and discussing your projects. Topics include:
|
| 10-14 Feb 2017 |
Phenome 2017 Venue: Hilton El Conquistador Resort, Tucson, Arizona, USA The Phenome 2017 conference is an important step in the development of a path toward training and collaboration among disciplines that are poised to address critical social issues and to generate greater understanding about plants and climate change. This first conference will bring together a multidisciplinary community comprising plant biologists, ecologists, engineers, agronomists, and computer scientists. Please share this flyer with your members of your department and encourage your students to register to attend the meeting. Highly relevant abstracts submitted by November 1 may be considered to give a talk during one of the sessions. Please visit www.phenome2017.org to register and learn more about the meeting. |
| 13-17 Feb 2017 |
EMBO Practical Course on Metabolomics Bioinformatics for Life Scientists Venue: European Bioinformatics Institute (EMBL-EBI) - Training Room 2 - Wellcome Genome Campus, Hinxton, Cambridge, CB10 1SD, UK Application opens: Monday August 08 2016 Application deadline: Friday November 11 2016 Participation: Open application with selection Contact: Maria Bacadare Goitia Registration fee: £350 Overview This course will provide an overview of key issues that affect metabolomics studies, handling datasets and procedures for the analysis of metabolomics data using bioinformatics tools. It will be delivered using a mixture of lectures, computer-based practical sessions and interactive discussions. The course will provide a platform for discussion of the key questions and challenges in the field of metabolomics, from study design to metabolite identification. Audience This course is aimed at PhD students, post-docs and researchers with at least one year’s experience in the field of metabolomics who are seeking to improve their skills in metabolomics data analysis. Participants ideally must have working experience using R (including a basic understanding of the syntax and ability to manipulate objects). For more information, please visit https://www.ebi.ac.uk/training/events/2017/embo-practical-course-metabolomics-bioinformatics-life-scientists-3. |
| 20 Feb to 17 Mar
2017 |
Metabolomics Data Processing and Data Analysis Online Course An online course by the Birmingham Metabolomics Training Center, University of Birmingham, UK hosted by Futurelearn Venue: Birmingham Metabolomics Training Centre, University of Birmingham, Birmingham, UK This four-week online course will explore the tools and approaches that are used to process and analyse metabolomics data, we will investigate the challenges that are typically encountered in the analysis of metabolomics data and provide solutions to overcome these problems. The course will be delivered using a combination of short videos, articles, discussions, and online workshops with step-by-step instructions and test data sets. We will provide quizzes, polls and peer review exercises each week, so that you can review your learning throughout the course. The material will be delivered over a four week period, with an estimated learning time of four hours per week. If you do not have time to complete the course during the 4-week period you will retain access to the course material to revisit, as you are able. Registration fee: Early-bird £200, Standard £220 For further information and registration details, please visit http://www.birmingham.ac.uk/facilities/metabolomics-training-centre/courses/Metabolomics-Data-Processing-and-Data-Analysis.aspx or contact bmtc@contacts.bham.ac.uk. |
| 13-15 Mar 2017 |
CPSA Metabolomics 2017 Venue: The University of Florida Clinical & Translational Science Institute, Gainesville, Florida, USA Make plans to attend the 3rd Annual Metabolomics Symposium on Clinical & Pharmaceutical Solutions through Analysis (CPSA Metabolomics 2017). This unique event is highly interactive and dedicated to the needs of the clinic. The program features updated perspectives and experiences on clinical and pharmaceutical analysis. Imagination and stimulating discussion are central to each CPSA Metabolomics session and event. Goal The goal of CPSA Metabolomics is to provide in-depth review of innovative technology and industry practices through open discussion of industry-related issues and needs. This annual event is specifically geared to the needs of professionals attempting to keep pace with faster development times and technology marketing managers attempting to benchmark emerging trends. Program The CPSA Metabolomics symposia and roundtables are highly interactive events where scientists share their experiences and visions in a collegial setting. The program will highlight speakers and sessions that provide real-world experiences with new technologies and critical insights into current issues and future needs. Education and specialized training are the foundation of all CPSA events. Each session at CPSA Metabolomics will address the current industrial landscape and the global need to bring products to market faster. The program chairs will promote discussion and exchange of experiences, ideas, and visions so that current processes that involve analytical measurement can be benchmarked and future strategies may be developed. Format The CPSA Metabolomics symposia and roundtables feature a unique format to allow scientists to openly share comprehensive perspectives on industry-related issues and needs. First-hand experiences with specific applications and technologies are openly discussed. Lively discussions in a collegial setting are a hallmark of the meeting. For more information, visit http://www.cpsa-metabolomics.com/2017/index.shtml.
|
| 11-13 May 2017 |
Metabolism in Time and Space: Emerging Links to Cellular and Developmental Programs Venue: EMBL Heidelberg, Germany T. Alexandrov, A. Aulehla, P. Dorrestein, O. Leyser, S. McKnight, N. Perrimon Deadlines
Topics
Latest News
Stay up to date! Add this event
easily to your calendar by
downloading the
iCal>>
Add
to calendar For more information, visit http://www.embo-embl-symposia.org/symposia/2017/EES17-01/index.html
|
| 17-18 May 2017 |
Conference on Food and Nutritional Metabolomics for Health Venue: The Ohio State University, Columbus, OH, USA The purpose of this two-day event is to disseminate state-of-the-art knowledge in the field of food and nutritional metabolomics and foster networking and collaboration among colleagues and industry partners. For more information please visit: https://discovery.osu.edu/focus-areas/foods-for-health/events/conference-2017.html. |
| 29 May to 2 June
2017 |
Workflow4Experimenters International Course (W4E2017) Venue: Paris, France Analyze your own data with Galaxy and the Workflow4metabolomics e-infrastructure! The next Workflow4Experimenters international course (W4E2017) will take place in Paris (May 29 to June 2, 2017). During this one-week course (entirely in English), you will learn how to use Galaxy and the W4M infrastructure to analyze your own LC-MS, GC-MS, or NMR data set. Morning sessions will be dedicated to methodology and tools. Afternoon sessions will be devoted to tutoring. Invited speakers: Tim Ebbels (Imperial College), Steffen Neumann (IPB Halle), and Ralf Weber (Birmingham University). Registrations: http://workflow4metabolomics.org Contact: contact@workflow4metabolomics.org |
| 26-29 June 2017 |
Metabolomics 2017 Venue: Brisbane, Australia  It is our pleasure to invite you to the 13th International Conference of the Metabolomics Society from 25-29th June, 2017 at the Brisbane Conference and Exhibition Centre (BCEC) in Brisbane, Australia. Brisbane is a vibrant, friendly, lifestyle city—home to leading medical research and a thriving industry hub, located in the heart of Australia’s premier tourist region. The BCEC is rated among the top three convention centres in the world, and was the venue of the 2014 G20 Leaders Summit. It is ideally located in the unique riverside cultural and lifestyle precinct at South Bank, which is an inner city oasis with riverfront parkland, rainforest pockets and Australia’s only city-based sand and swimming beach as well as Australia’s newest and largest Gallery of Modern Art, cafes, restaurants and stylish shops. The conference has the theme of Building Bridges and under this banner extends its reach to the systems biology / genome-scale modelling community, as well as to the analytical chemistry / natural products chemistry community. In addition, the program features thematic streams for advancing the field, for food and environmental metabolomics, and for health and wellness. In addition, a deeper engagement between researchers within the Asia Pacific region is a natural focus for a conference held in Brisbane to promote metabolomics research, build and strengthen networks in the region. We invite you to attend an exciting scientific program comprising 27 oral sessions, 5 plenaries, 4 poster sessions, sponsored luncheons, as well as several keynote lectures and workshops. We will continue the successful tradition of satellite workshops to the conference in the afternoon of Sunday 25th June and the morning of Monday 26th June. Additionally, we have planned a range of social activities, including a welcome reception, an early-career researcher mixer and a conference dinner in the iconic BCEC Plaza Ballroom to give you a true Aussie-style experience. Brisbane is the ideal opportunity for delegates to enjoy a microcosm of Australia’s iconic experiences. World heritage listed rainforests, amazing beaches, islands, wineries and the internationally famous Australia Zoo—home of the crocodile hunter—are all easily accessible within an hour of the city. You can even do day trips to the Barrier Reef from Brisbane. On behalf of the Local Organising Committee and the Metabolomics Society Board we are excited to once again invite you to Metabolomics 2017—we are looking forward to welcoming you down under! Prof. Melissa Fitzgerald, School of Agriculture and Food Sciences & Dr. Horst Joachim Schirra, Centre for Advanced Imaging, The University of Queensland, Brisbane, Australia For more information, visit http://metabolomics2017.org/. |
Metabolomics Jobs |
This is a resource for
advertising positions in metabolomics. If
you have a job you would like posted in this
newsletter, please email Ian Forsythe (metabolomics.innovation@gmail.com).
Job postings will be carried for a maximum
of four issues (eight weeks) unless the
position is filled prior to that date.
Jobs Offered
| Job Title | Employer | Location | Posted | Closes | Source |
|---|---|---|---|---|---|
| Research Fellow in Mass
Spectrometry Metabolomics - 55220 |
University of Birmingham | Birmingham, UK | 26-Dec-2016 | 22-Jan-2017 |
University
of Birmingham |
| UCD Post-doctoral
Research Fellow Level 2 |
University College Dublin | Dublin,
Ireland |
22-Dec-2016 | 9-Jan-2017 |
University
College Dublin |
| Postdoctoral Fellowship
in Metabolomics |
Lund University Diabetes Centre | Malmö, Sweden | 21-Dec-2016 | |
Lund University Diabetes Centre |
| Post-Doctoral Research
Fellow
(Bioinformatician/Biostatistician) |
Edith Cowan University | Perth, Australia | 3-Dec-2016 | 22-Jan-2017 |
Edith
Cowan
University |
| Post-Doctoral Research
Fellow (Biochemist/Analytical Chemist) |
Edith Cowan University | Perth, Australia | 3-Dec-2016 | 22-Jan-2017 | Edith Cowan University |
| Field Application Specialist Metabolomics-Mass Spectrometry | BIOCRATES Life Sciences AG | East Coast, USA | 25-Nov-2016 |
|
BIOCRATES
Life Sciences
AG |
| Postdoctoral Researcher
in the functional metabolomics of
immuno-metabolism |
Karolinska Institutet | Stockholm,
Sweden |
24-Nov-2016 | 31-Jan-2017 | Karolinska Institutet |
| Postdoctoral Researcher
in the functional metabolomics of
lipid mediators in chronic respiratory
disease |
Karolinska Institutet | Stockholm, Sweden | 14-Nov-2016 | 2-Jan-2017 |
Nature
Jobs |
| PhD Project "Minimal
invasive biomarkers for the
personalized treatment of neonatal
Hypoxic-Ischemic Encephalopathy (HIE)" |
Health Research Institute of Hospital La Fe | Valencia,
Spain |
10-Nov-2016 |
|
La Caixa and the European Union; search for "Minimal invasive biomarkers for the personalized treatment of neonatal Hypoxic-Ischemic Encephalopathy (HIE)" |
| Post-doctoral
Fellow/Staff Scientist
[Musculoskeletal Metabolomics] |
Oklahoma Medical Research Foundation | Oklahoma City, Oklahoma, USA | 7-Nov-2016 | Until filled | Metabolomics Society |
|
Ian J. Forsythe Double AB, MSc Editor, MetaboNews Department
of Computing Science
University of Alberta 221 Athabasca Hall Edmonton, AB, T6G 2E8, Canada Email: metabolomics.innovation@gmail.com Website: http://www.metabonews.ca LinkedIn: https://ca.linkedin.com/in/ian-forsythe-465512128 Twitter: http://twitter.com/MetaboNews Google+: https://plus.google.com/118323357793551595134 Facebook: http://www.facebook.com/metabonews |
This newsletter is
published in partnership between The
Metabolomics Innovation Centre
(TMIC, http://www.metabolomicscentre.ca/) and
the Metabolomics Society (http://www.metabolomicssociety.org)
for the benefit of the worldwide
metabolomics community.
|