MxP® Heart Score—a
metabolomics-based biomarker for the early detection of
heart failure with reduced ejection fraction (HFrEF)
"The new heart failure biomarker we jointly validated with
Metanomics GmbH/Metanomics Health GmbH has the potential to
improve the management of this devastating disease
significantly. It is showing superior performance as compared
to a reference biomarker currently used to evaluate patients
already showing symptoms. Of note, the newly identified
biomarker may also be useful in patients with early stages of
the disease."
Prof. Dr. Hugo Katus, Head of the Cardiology Department of
Ruprecht-Karls-University
Feature article contributed by Dr.
Philipp Schatz, Head of Biomarker Program, Metanomics
Health GmbH, Tegeler Weg 33, 10589 Berlin, Germany
In the Western world, an aging population, a sedentary
lifestyle, and a rising number of cardiac disease comorbidities
are leading to an increasing prevalence of congestive heart
failure (CHF accounts for 5% of all hospital admissions) and
present a heavy burden to health care systems. In particular,
subjects over the age of 65 with additional risk factors, such
as elevated arterial blood pressure, coronary artery disease,
and/or type 2 diabetes have a high risk of developing CHF
(prevalence up to 20%)
1,2,3.
Since CHF develops long before the onset of signs and symptoms,
accurate early detection and, in the case of HFrEF,
pharmacological and behavioral interventions are of the utmost
importance. However, screening tools for left ventricular
systolic dysfunction available to the primary care physician
suffer from unacceptably high false-positive rates and lead to a
high number of patients being recommended for costly follow-up
procedures. Thus, HFrEF screening programs have not yet been
established.
Researchers from the University Clinics of Heidelberg, Berlin,
Kiel, and from Metanomics GmbH/ Metanomics Health GmbH, Berlin
have identified and validated a new biomarker for the improved
management of this common, costly, disabling, and potentially
deadly condition by enrolling and analyzing more than 800 CHF
patients in different disease stages together with 300 healthy
controls (
Figure
1). A metabolite-based biomarker on top of NT-proBNP for
the detection of HFrEF was identified and validated. The new
biomarker is capable of detecting left ventricular systolic
dysfunction with high precision, particularly in the early
disease stages. This new diagnostic biomarker shows a
performance superior to the reference marker NT-proBNP alone (at
a comparable negative predictive value, the positive predictive
value for the new marker almost doubles,
Table
1) and has in the meantime been developed into an LC-MS/MS
CLIA-ready assay.
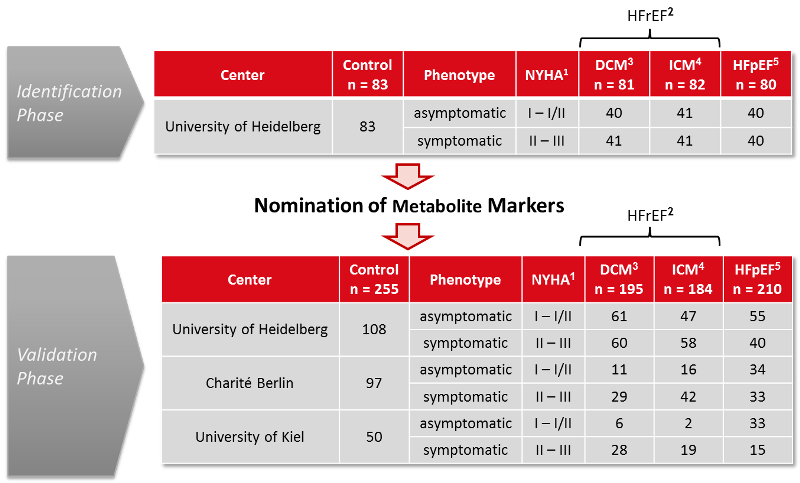 Figure 1. Study Design – flow chart
Figure 1. Study Design – flow chart
1NYHA: Classification according to the New York
Heart Association,
2HFrEF: Heart failure with
reduced ejection fraction,
3DCM: Dilated
cardiomyopathy,
4ICM: Ischemic cardiomyopathy, and
5HFpEF:
Heart failure with preserved ejection fraction.
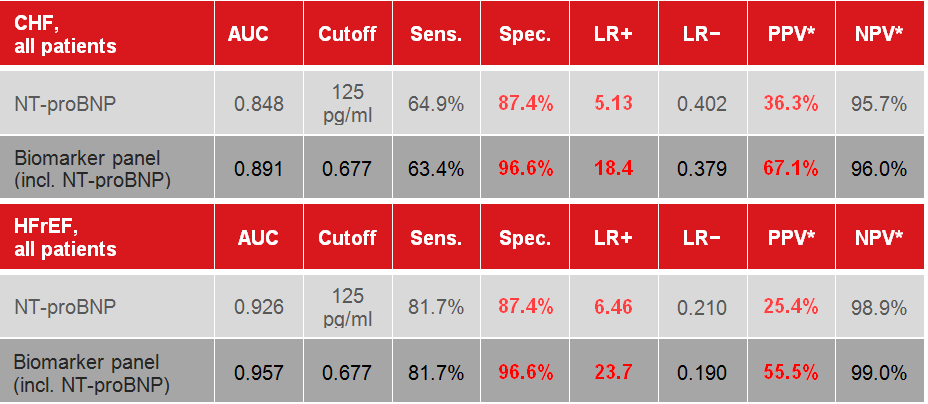 Table 1.
Table 1. Clinical performance of the CLIA-ready
metabolite-based early detection biomarker in patients with
heart failure in general (CHF, all patients)
4
and in patients with systolic dysfunction (HFrEF, all patients)
5).
Abbreviations: AUC, area under the curve of the receiver
operating characteristic (ROC); Sens., sensitivity; Spec.,
specificity; LR+, positive likelihood ratio; LR−, negative
likelihood ratio; NPV, negative predictive value; PPV, positive
predictive value. *Prevalence of heart failure in the
Medicare-eligible population of the US was 9-12% between 1994
and 2003 and is assumed here for the purpose of estimating PPV
and NPV with 10%, half of which are affected by HFrEF
1,2.
The new HFrEF screening biomarker indicates alterations in
numerous metabolic pathways, involving lipids and amino acids in
CHF patients. The most consistent alterations across centres
were observed for lipid classes including complex lipids such as
sphingolipids. In addition to the significant clinical value of
early HFrEF diagnosis, the combination of several metabolites
with the commercially available NT-proBNP test has the potential
to overcome the limitations of single feature markers in a
complex disease such as HFrEF. The new diagnostic biomarker
reflects multiple pathology-associated metabolic changes in
addition to the results achieved by the commercially available
NT-proBNP test, and is thus more robust in a diverse group of
subjects, many of which suffer from comorbidities.
For routine CLIA-ready testing, a diagnostic assay has been
developed in parallel. This prototype assay is based on an
optimized LC-MS/MS technology for selected analytes. The new
CLIA-ready assay was developed using samples from the original
validation cohort, split into a training set and a test set
comprised of 410 and 219 patients, respectively.
Figure
2 shows NT-proBNP concentrations and the biomarker
prediction scores for the new metabolite-based marker for all
HFrEF subjects and healthy controls in the combined training and
test set. A cutoff value of 0.677 was chosen for the biomarker
prediction score to match the sensitivity with that of the
NT-proBNP test alone with respect to the detection of HFrEF in
the training set. As shown in
Figure
2, the new metabolite-based CLIA-ready assay resulted in
fewer healthy controls (blue) being misclassified as diseased
compared with NT-proBNP alone. This increase in specificity at
comparable sensitivity results in a strongly increased
diagnostic likelihood ratio and PPV, rendering the new
metabolite-based biomarker suitable for heart failure screening
programs.
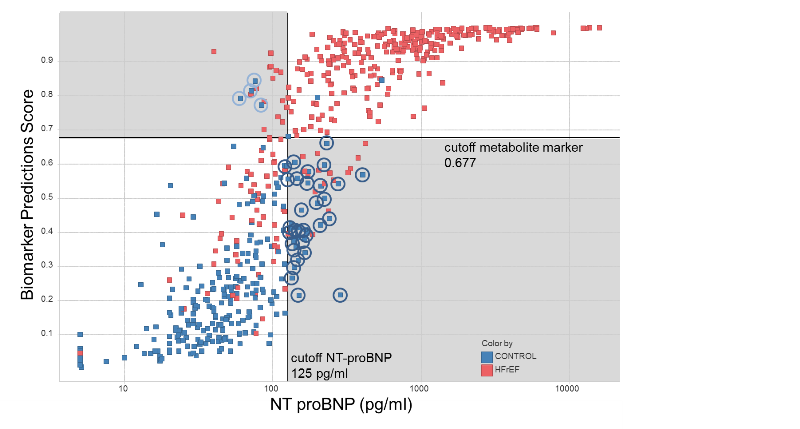 Figure 2.
Figure 2. Scatter plot of the prediction score of the
new metabolite-based HFrEF marker + NT-proBNP vs. the plasma
concentration of NT-proBNP alone in subjects with reduced
systolic function (LVEF < 50%) and in healthy controls. Solid
lines represent the cutoff values for NT-proBNP (125 pg/ml,
vertical line) and the new metabolite-based biomarker
(prediction score 0.677, horizontal line). Dark blue circled
control subjects are misclassified by NT-proBNP and are
correctly identified using the metabolic test. Light blue
circled control subjects are false positives using the metabolic
test and are correctly identified as healthy by NT proBNP.
The new CLIA-ready assay generates quantitative results.
Positive and negative controls are prepared by lyophilization of
different amounts of plasma to meet the positive and negative
cutoffs for all metabolites. Samples for the daily quality
control of the instrument performance are prepared by extracting
commercially available plasma with extraction solvent. The
improvement in diagnostic performance of the new CLIA-ready
assay over NT-proBNP only is shown in
Table
1. Details of the methods will soon be published in a
peer-reviewed publication.
Data were derived from a large heart failure cohort study
conducted between 2008 and 2013. The multicenter trial was
co-funded by the German Network for Genome Research (NGFN).
Currently a peer review publication is being prepared.
Further details of the MxP
® Heart Score biomarker
were presented at the European Society of Cardiology (ESC)
Meeting in Amsterdam on September 4
th, 2013.
Metanomics Health is currently seeking academic and diagnostic
partners to further develop and market its HFrEF screening
biomarker.
Metanomics Health, a BASF Group company, is the world-leading
company offering targeted and non-targeted metabolomics to
healthcare, nutrition, and bioprocessing partners in industry
and academia. In parallel with the service business, Metanomics
Health is funding and conducting a comprehensive clinical
biomarker program, addressing a wide range of questions with
high unmet medical needs.
Find out more about Metanomics Health GmbH at www.metanomics-health.com
References
- Bertoni AG, Hundley WG, Massing MW, et al. (2004).
Diabetes Care 27(3):699-703.
- McMurray JJ, Pfeffer MA (2005). "Heart failure". Lancet
365(9474):1877-89.
- Dickstein K, Cohen-Solal A, Filippatos G, et al.
(2008). Eur J Heart Fail. 10(10):933-89.
- CHF all patients, left ventricle wall thickness > 55 mm
AND LVEF <50% OR >50% stenosis AND LVEF <50% OR
Cardiac septum > 11 mm AND posterior wall thickness >
11, all patients.
- Left Ventricular Ejection Fraction ≤ 50%.
Please note: If you know of any metabolomics research
programs, software, databases, statistical methods,
meetings, workshops, or training sessions that we should
feature in future issues of this newsletter, please email
Ian Forsythe at metabolomics.innovation@gmail.com.









